The potential for AI-powered Shiny app prototyping with Shiny Assistant

Get our email updates
Interested in learning more about Posit + AI tools? Join our email list.
The pharmaceutical industry deals with complex datasets, from clinical trial results to genomic analyses. Shiny apps offer an interactive way to explore these datasets, communicate findings, and support data-driven decisions. The use of Shiny in life sciences has seen tremendous growth across the spectrum of drug development, from evaluating novel disease state targets to interactive submission packages. However, developing a Shiny app from scratch can be time-intensive, requiring both domain expertise and technical skills. Generative AI can make it faster and easier to create interactive tools and bridge the gap between ideas and implementation.
Announced during posit::conf(2024), Shiny Assistant is a chatbot-powered AI tool that guides users in creating Shiny apps. Users describe app requirements in plain language and receive tailored code suggestions. They can then refine prototypes in real time while leveraging the latest innovations from the Shiny team.
For example, Shiny Assistant can quickly create a basic app structure for visualizing clinical trial data, saving hours that would otherwise be spent on boilerplate code or troubleshooting layout issues. This not only saves time but also offers a hands-on way for users to practice and improve their coding skills. Below is a walkthrough showcasing Shiny Assistant’s capabilities. Note that AI’s inherent limitations results in slight differences between the demonstration and an app generated during a live interaction.
How It Works
Describe the app
Begin by chatting with the AI. To verify that Shiny Assistant understood the pharma terms I planned to use, I started by asking, “Do you know what Adverse Event Analysis Datasets (ADAE) are?” It replied yes, it was familiar with ADAE. I verified the information to ensure it was correct.
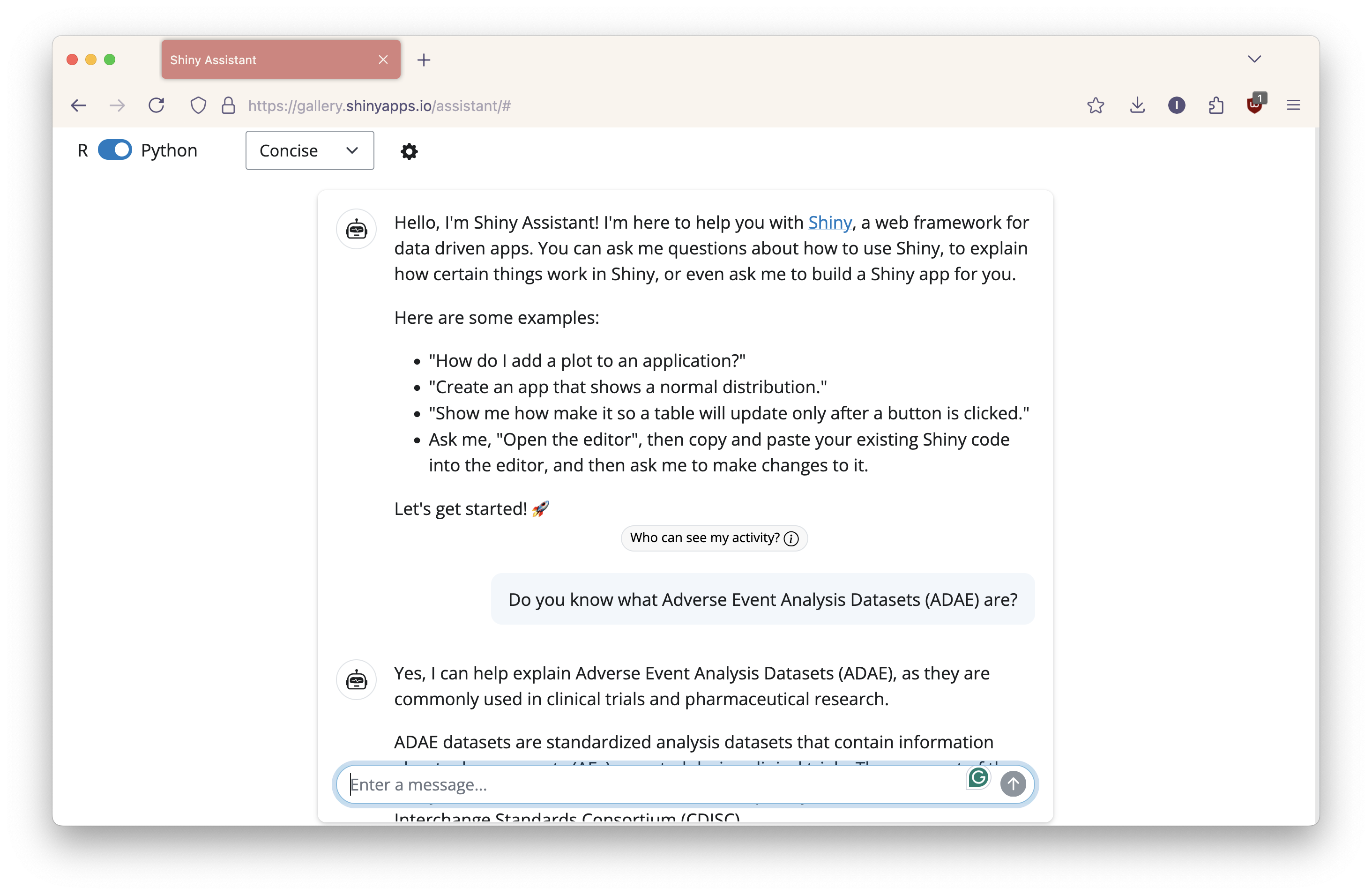
It then followed up with, “Would you like to see an example of a Shiny app that could be used to analyze ADAE data? I could create one that demonstrates common types of adverse event analyses,” to which I replied, “Yes, please”.
Chat history
Hello, I’m Shiny Assistant! I’m here to help you with Shiny, a web framework for data driven apps. You can ask me questions about how to use Shiny, to explain how certain things work in Shiny, or even ask me to build a Shiny app for you.
Here are some examples:
"How do I add a plot to an application?"
"Create an app that shows a normal distribution."
"Show me how make it so a table will update only after a button is clicked."
Ask me, "Open the editor", then copy and paste your existing Shiny code into the editor, and then ask me to make changes to it.Let’s get started! 🚀 Who can see my activity?
Do you know what Adverse Event Analysis Dataset (ADAE) are?
Let me explain what an Adverse Event Analysis Dataset (ADAE) is, as it’s commonly used in clinical trials and pharmaceutical research.
ADAE is a standardized dataset format that’s part of the Study Data Tabulation Model (SDTM) and Analysis Data Model (ADaM) used in clinical trials. It contains detailed information about adverse events (unwanted medical occurrences) that participants experience during a clinical study.
Key variables typically found in ADAE datasets include:
Subject identifiers
Treatment/study arm information
Adverse event terms (usually coded using MedDRA terminology)
Start and end dates of adverse events
Severity/intensity
Seriousness
Action taken
Relationship to study drug
Outcome
Duration
Time to onsetCommon analyses performed on ADAE datasets include:
Frequency counts of adverse events
Time-to-event analyses
Severity distributions
Relationship to study drug assessments
Comparative analyses between treatment groupsThis type of dataset is crucial for safety monitoring in clinical trials and is often used to create safety tables and listings for regulatory submissions and clinical study reports.
Would you like to see an example of a Shiny app that could be used to analyze ADAE data? I could create one that demonstrates common types of adverse event analyses.
Yes, please.
Receive code suggestions
This is where the magic of Shiny Assistant happens. Once I ask it to build an app, the chatbot generates an app.R file that includes UI elements and server logic. The UI is organized using bslib’s card layout, featuring a sidebar for analysis selection and filtering options, and a main panel displaying the selected analysis results.
On the right-hand side of the Assistant, a new window opens. Powered by Shinylive, it runs the code provided in the conversation window and builds the application directly in the browser. This allows you to view the dashboard, edit the code, and rerun the application without needing to install R, an IDE, or any packages locally.
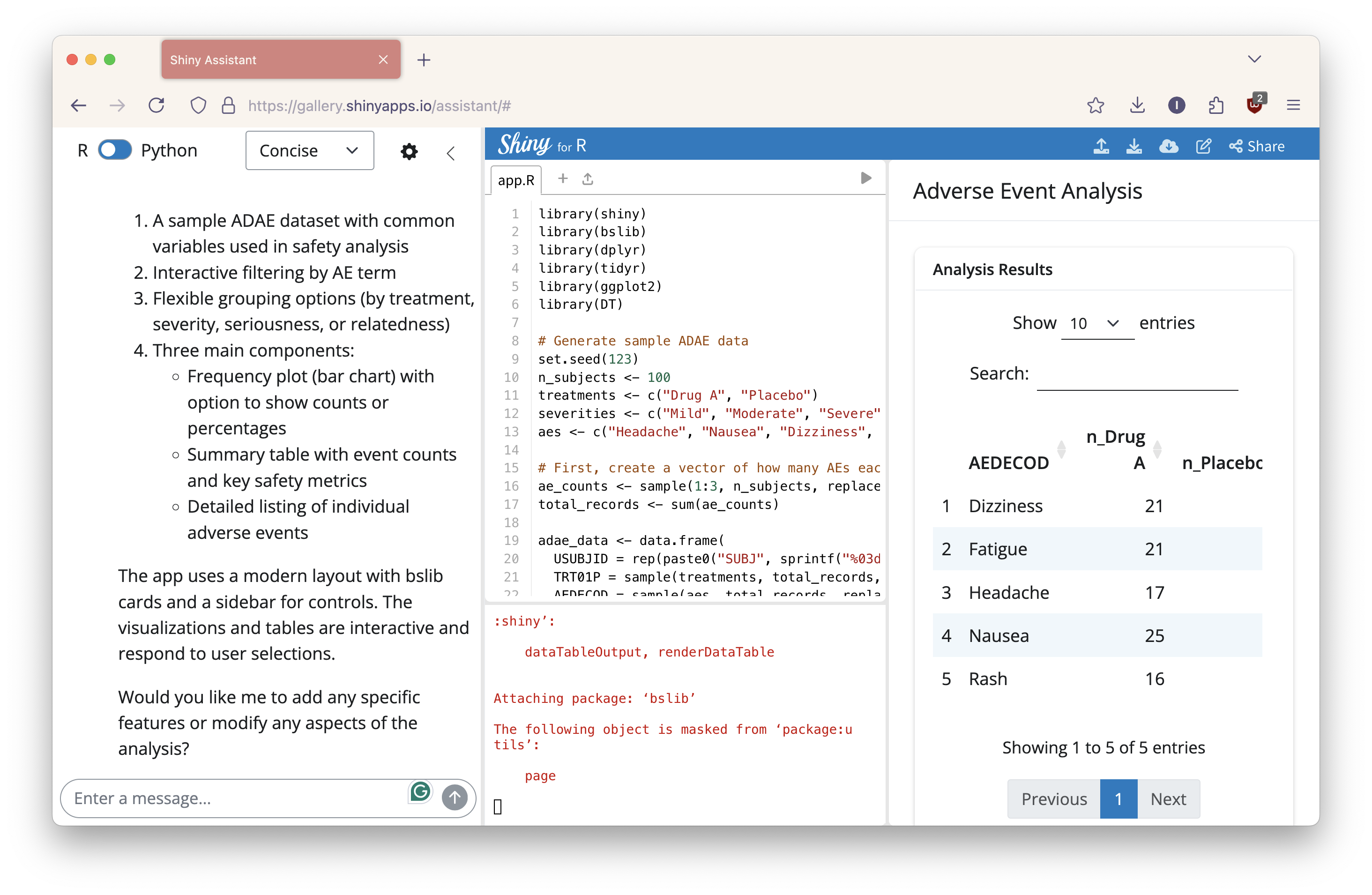
The first time I ran this, the Shiny Assistant produced code but encountered an error: Error: arguments imply differing number of rows: 202, 200. I informed the Assistant, and it explained the issue before regenerating the code. This time, the Shiny app successfully ran in the Shinylive viewer.
The resulting app featured a summary table, a bar plot showing severity by treatment, and a density plot for time to onset. While it required a couple iterations to resolve the initial issue, the Shiny Assistant provided a compelling prototype early on to jumpstart the development process.
Chat history
I’ll create a Shiny app that demonstrates common analyses for adverse event data. This app will include features like summary tables of adverse events by severity and treatment group, visualizations of adverse event frequencies, and time-to-onset analysis.
app.R
library(shiny)
library(bslib)
library(dplyr)
library(tidyr)
library(ggplot2)
library(DT)
# Generate sample ADAE data
set.seed(123)
n_subjects <- 100
treatments <- c("Drug A", "Placebo")
severities <- c("Mild", "Moderate", "Severe")
aes <- c("Headache", "Nausea", "Dizziness", "Fatigue", "Rash")
adae_data <- data.frame(
USUBJID = rep(paste0("SUBJ", sprintf("%03d", 1:n_subjects)),
sample(1:3, n_subjects, replace = TRUE)) %>% unlist(),
TRT01P = sample(treatments, n_subjects * 2, replace = TRUE),
AEDECOD = sample(aes, n_subjects * 2, replace = TRUE),
AESEV = sample(severities, n_subjects * 2, replace = TRUE,
prob = c(0.6, 0.3, 0.1)),
ASTDY = sample(1:100, n_subjects * 2, replace = TRUE),
AESER = sample(c("Y", "N"), n_subjects * 2, replace = TRUE,
prob = c(0.1, 0.9))
)
ui <- page_sidebar(
title = "Adverse Event Analysis",
sidebar = sidebar(
selectInput("analysis_type", "Select Analysis",
choices = c("AE Summary Table",
"AE by Severity",
"Time to Onset Analysis")),
checkboxGroupInput("severity_filter", "Filter by Severity",
choices = severities,
selected = severities),
checkboxInput("serious_only", "Show Only Serious AEs", FALSE)
),
# Main panel with cards
card(
card_header("Analysis Results"),
card_body(
conditionalPanel(
condition = "input.analysis_type == 'AE Summary Table'",
DTOutput("ae_table")
),
conditionalPanel(
condition = "input.analysis_type == 'AE by Severity'",
plotOutput("severity_plot")
),
conditionalPanel(
condition = "input.analysis_type == 'Time to Onset Analysis'",
plotOutput("onset_plot")
)
)
)
)
server <- function(input, output) {
# Filtered dataset based on user inputs
filtered_data <- reactive({
data <- adae_data
if (!is.null(input$severity_filter)) {
data <- data %>% filter(AESEV %in% input$severity_filter)
}
if (input$serious_only) {
data <- data %>% filter(AESER == "Y")
}
data
})
# AE Summary Table
output$ae_table <- renderDT({
filtered_data() %>%
group_by(AEDECOD, TRT01P) %>%
summarise(
n = n(),
n_serious = sum(AESER == "Y"),
.groups = "drop"
) %>%
pivot_wider(
names_from = TRT01P,
values_from = c(n, n_serious),
values_fill = 0
) %>%
datatable(options = list(pageLength = 10))
})
# AE by Severity Plot
output$severity_plot <- renderPlot({
filtered_data() %>%
ggplot(aes(x = AEDECOD, fill = AESEV)) +
geom_bar(position = "dodge") +
facet_wrap(~TRT01P) +
theme_minimal() +
theme(axis.text.x = element_text(angle = 45, hjust = 1)) +
labs(x = "Adverse Event", y = "Count", fill = "Severity") +
scale_fill_brewer(palette = "YlOrRd")
})
# Time to Onset Plot
output$onset_plot <- renderPlot({
filtered_data() %>%
ggplot(aes(x = ASTDY, fill = TRT01P)) +
geom_density(alpha = 0.5) +
facet_wrap(~AEDECOD) +
theme_minimal() +
labs(x = "Study Day of Onset", y = "Density", fill = "Treatment") +
scale_fill_brewer(palette = "Set2")
})
}
shinyApp(ui, server)This Shiny app provides three common types of adverse event analyses:
AE Summary Table: Shows the frequency of each adverse event by treatment group, including counts of serious events.
AE by Severity: Visualizes the distribution of adverse events by severity level and treatment group.
Time to Onset Analysis: Shows the density distribution of when adverse events occurred during the study.The app includes filters for:
Severity levels (Mild, Moderate, Severe)
Serious events onlyThe sample data is generated within the app for demonstration purposes, but in practice, you would typically load your ADAE dataset from an external source.
The UI is organized using bslib’s card layout, with a sidebar for analysis selection and filtering options. The main panel displays the selected analysis results.
Would you like me to add any specific features or modify any aspects of this analysis?
There is an error in your code: Error: arguments imply differing number of rows: 202, 200
I understand the issue. The error occurs because we’re using n_subjects * 2 in some places and the rep() function with sample(1:3, n_subjects, replace = TRUE) in another, which creates vectors of different lengths. Let’s fix this by making the data generation more consistent.
app.R
library(shiny)
library(bslib)
library(dplyr)
library(tidyr)
library(ggplot2)
library(DT)
# Generate sample ADAE data
set.seed(123)
n_subjects <- 100
treatments <- c("Drug A", "Placebo")
severities <- c("Mild", "Moderate", "Severe")
aes <- c("Headache", "Nausea", "Dizziness", "Fatigue", "Rash")
# First, create a vector of how many AEs each subject will have
ae_counts <- sample(1:3, n_subjects, replace = TRUE)
total_records <- sum(ae_counts)
adae_data <- data.frame(
USUBJID = rep(paste0("SUBJ", sprintf("%03d", 1:n_subjects)), ae_counts),
TRT01P = sample(treatments, total_records, replace = TRUE),
AEDECOD = sample(aes, total_records, replace = TRUE),
AESEV = sample(severities, total_records, replace = TRUE,
prob = c(0.6, 0.3, 0.1)),
ASTDY = sample(1:100, total_records, replace = TRUE),
AESER = sample(c("Y", "N"), total_records, replace = TRUE,
prob = c(0.1, 0.9))
)
ui <- page_sidebar(
title = "Adverse Event Analysis",
sidebar = sidebar(
selectInput("analysis_type", "Select Analysis",
choices = c("AE Summary Table",
"AE by Severity",
"Time to Onset Analysis")),
checkboxGroupInput("severity_filter", "Filter by Severity",
choices = severities,
selected = severities),
checkboxInput("serious_only", "Show Only Serious AEs", FALSE)
),
# Main panel with cards
card(
card_header("Analysis Results"),
card_body(
conditionalPanel(
condition = "input.analysis_type == 'AE Summary Table'",
DTOutput("ae_table")
),
conditionalPanel(
condition = "input.analysis_type == 'AE by Severity'",
plotOutput("severity_plot")
),
conditionalPanel(
condition = "input.analysis_type == 'Time to Onset Analysis'",
plotOutput("onset_plot")
)
)
)
)
server <- function(input, output) {
# Filtered dataset based on user inputs
filtered_data <- reactive({
data <- adae_data
if (!is.null(input$severity_filter)) {
data <- data %>% filter(AESEV %in% input$severity_filter)
}
if (input$serious_only) {
data <- data %>% filter(AESER == "Y")
}
data
})
# AE Summary Table
output$ae_table <- renderDT({
filtered_data() %>%
group_by(AEDECOD, TRT01P) %>%
summarise(
n = n(),
n_serious = sum(AESER == "Y"),
.groups = "drop"
) %>%
pivot_wider(
names_from = TRT01P,
values_from = c(n, n_serious),
values_fill = 0
) %>%
datatable(options = list(pageLength = 10))
})
# AE by Severity Plot
output$severity_plot <- renderPlot({
filtered_data() %>%
ggplot(aes(x = AEDECOD, fill = AESEV)) +
geom_bar(position = "dodge") +
facet_wrap(~TRT01P) +
theme_minimal() +
theme(axis.text.x = element_text(angle = 45, hjust = 1)) +
labs(x = "Adverse Event", y = "Count", fill = "Severity") +
scale_fill_brewer(palette = "YlOrRd")
})
# Time to Onset Plot
output$onset_plot <- renderPlot({
filtered_data() %>%
ggplot(aes(x = ASTDY, fill = TRT01P)) +
geom_density(alpha = 0.5) +
facet_wrap(~AEDECOD) +
theme_minimal() +
labs(x = "Study Day of Onset", y = "Density", fill = "Treatment") +
scale_fill_brewer(palette = "Set2")
})
}
shinyApp(ui, server)I fixed the data generation part of the code. The main changes are:
First calculate how many AEs each subject will have using ae_counts
Calculate the total number of records needed using total_records <- sum(ae_counts)
Use this total_records value consistently when generating all columns of the datasetIterate and refine
From here, the user can continue the conversation to refine the code, add new features, or enhance interactivity. For example, Shiny Assistant provided sample code for its app demonstration. However, I wanted to use my own ADAE dataset (generated with the random.cdisc.data package). Since my dataset’s variable names differed from those in Shiny Assistant’s code, I asked it to update the app accordingly: “Can you replace variable names throughout the app? For the treatments variable, use TRT01A; for severity, use AESEV; and for aes, use AESEQ.” Shiny Assistant promptly updated the code to match my dataset.
Next, I requested, “Can you add some compelling visualizations?” In response, the Shiny Assistant introduced a heatmap, a forest plot, and enhanced the existing visualizations.
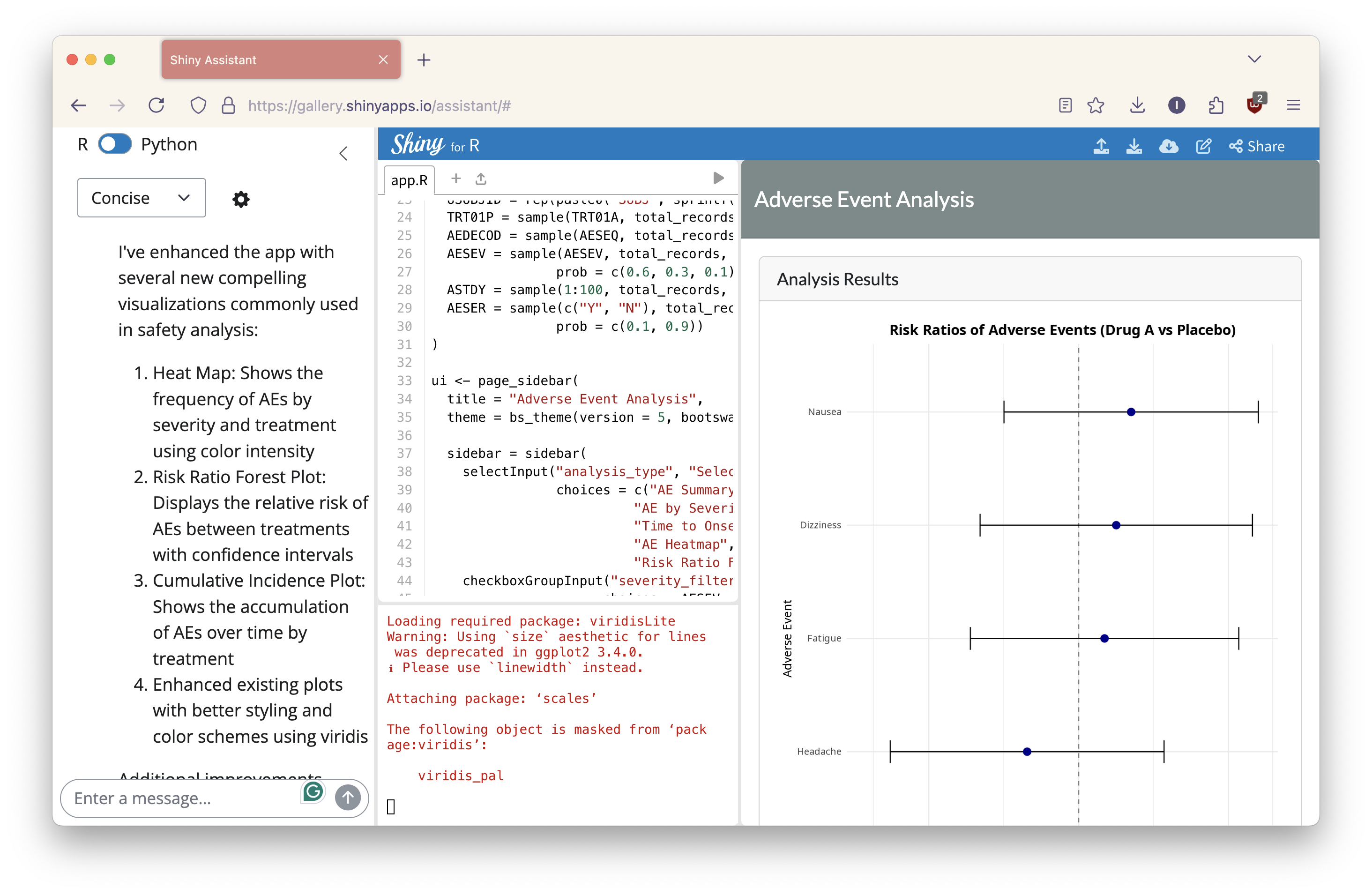
Chat history
Can you replace variable names throughout the app? For the treatments variable, can you instead use TRT01A, for severity, use AESEV instead, and for aes, use AESEQ.
I’ll help you replace those variable names throughout the app. Here’s the modified version:
app.R
library(shiny)
library(bslib)
library(dplyr)
library(tidyr)
library(ggplot2)
library(DT)
# Generate sample ADAE data
set.seed(123)
n_subjects <- 100
TRT01A <- c("Drug A", "Placebo")
AESEV <- c("Mild", "Moderate", "Severe")
AESEQ <- c("Headache", "Nausea", "Dizziness", "Fatigue", "Rash")
# First, create a vector of how many AEs each subject will have
ae_counts <- sample(1:3, n_subjects, replace = TRUE)
total_records <- sum(ae_counts)
adae_data <- data.frame(
USUBJID = rep(paste0("SUBJ", sprintf("%03d", 1:n_subjects)), ae_counts),
TRT01P = sample(TRT01A, total_records, replace = TRUE),
AEDECOD = sample(AESEQ, total_records, replace = TRUE),
AESEV = sample(AESEV, total_records, replace = TRUE,
prob = c(0.6, 0.3, 0.1)),
ASTDY = sample(1:100, total_records, replace = TRUE),
AESER = sample(c("Y", "N"), total_records, replace = TRUE,
prob = c(0.1, 0.9))
)
ui <- page_sidebar(
title = "Adverse Event Analysis",
sidebar = sidebar(
selectInput("analysis_type", "Select Analysis",
choices = c("AE Summary Table",
"AE by Severity",
"Time to Onset Analysis")),
checkboxGroupInput("severity_filter", "Filter by Severity",
choices = AESEV,
selected = AESEV),
checkboxInput("serious_only", "Show Only Serious AEs", FALSE)
),
# Main panel with cards
card(
card_header("Analysis Results"),
card_body(
conditionalPanel(
condition = "input.analysis_type == 'AE Summary Table'",
DTOutput("ae_table")
),
conditionalPanel(
condition = "input.analysis_type == 'AE by Severity'",
plotOutput("severity_plot")
),
conditionalPanel(
condition = "input.analysis_type == 'Time to Onset Analysis'",
plotOutput("onset_plot")
)
)
)
)
server <- function(input, output) {
# Filtered dataset based on user inputs
filtered_data <- reactive({
data <- adae_data
if (!is.null(input$severity_filter)) {
data <- data %>% filter(AESEV %in% input$severity_filter)
}
if (input$serious_only) {
data <- data %>% filter(AESER == "Y")
}
data
})
# AE Summary Table
output$ae_table <- renderDT({
filtered_data() %>%
group_by(AEDECOD, TRT01P) %>%
summarise(
n = n(),
n_serious = sum(AESER == "Y"),
.groups = "drop"
) %>%
pivot_wider(
names_from = TRT01P,
values_from = c(n, n_serious),
values_fill = 0
) %>%
datatable(options = list(pageLength = 10))
})
# AE by Severity Plot
output$severity_plot <- renderPlot({
filtered_data() %>%
ggplot(aes(x = AEDECOD, fill = AESEV)) +
geom_bar(position = "dodge") +
facet_wrap(~TRT01P) +
theme_minimal() +
theme(axis.text.x = element_text(angle = 45, hjust = 1)) +
labs(x = "Adverse Event", y = "Count", fill = "Severity") +
scale_fill_brewer(palette = "YlOrRd")
})
# Time to Onset Plot
output$onset_plot <- renderPlot({
filtered_data() %>%
ggplot(aes(x = ASTDY, fill = TRT01P)) +
geom_density(alpha = 0.5) +
facet_wrap(~AEDECOD) +
theme_minimal() +
labs(x = "Study Day of Onset", y = "Density", fill = "Treatment") +
scale_fill_brewer(palette = "Set2")
})
}
shinyApp(ui, server)I’ve made the following changes:
Changed treatments to TRT01A
Changed severities to AESEV
Changed aes to AESEQThe functionality of the app remains the same, only the variable names have been updated to match the requested naming convention.
Can you add some compelling visualizations?
I’ll enhance the app with additional, more compelling visualizations. I’ll add a heatmap, a forest plot for risk ratios, and improve the existing plots.
app.R
library(shiny)
library(bslib)
library(dplyr)
library(tidyr)
library(ggplot2)
library(DT)
library(viridis)
library(forestplot)
library(scales)
# Generate sample ADAE data
set.seed(123)
n_subjects <- 100
TRT01A <- c("Drug A", "Placebo")
AESEV <- c("Mild", "Moderate", "Severe")
AESEQ <- c("Headache", "Nausea", "Dizziness", "Fatigue", "Rash")
# First, create a vector of how many AEs each subject will have
ae_counts <- sample(1:3, n_subjects, replace = TRUE)
total_records <- sum(ae_counts)
adae_data <- data.frame(
USUBJID = rep(paste0("SUBJ", sprintf("%03d", 1:n_subjects)), ae_counts),
TRT01P = sample(TRT01A, total_records, replace = TRUE),
AEDECOD = sample(AESEQ, total_records, replace = TRUE),
AESEV = sample(AESEV, total_records, replace = TRUE,
prob = c(0.6, 0.3, 0.1)),
ASTDY = sample(1:100, total_records, replace = TRUE),
AESER = sample(c("Y", "N"), total_records, replace = TRUE,
prob = c(0.1, 0.9))
)
ui <- page_sidebar(
title = "Adverse Event Analysis",
theme = bs_theme(version = 5, bootswatch = "flatly"),
sidebar = sidebar(
selectInput("analysis_type", "Select Analysis",
choices = c("AE Summary Table",
"AE by Severity",
"Time to Onset Analysis",
"AE Heatmap",
"Risk Ratio Forest Plot")),
checkboxGroupInput("severity_filter", "Filter by Severity",
choices = AESEV,
selected = AESEV),
checkboxInput("serious_only", "Show Only Serious AEs", FALSE)
),
# Main panel with cards
layout_columns(
col_widths = 12,
card(
card_header("Analysis Results"),
card_body(
conditionalPanel(
condition = "input.analysis_type == 'AE Summary Table'",
DTOutput("ae_table")
),
conditionalPanel(
condition = "input.analysis_type == 'AE by Severity'",
plotOutput("severity_plot", height = "600px")
),
conditionalPanel(
condition = "input.analysis_type == 'Time to Onset Analysis'",
plotOutput("onset_plot", height = "600px")
),
conditionalPanel(
condition = "input.analysis_type == 'AE Heatmap'",
plotOutput("heatmap_plot", height = "600px")
),
conditionalPanel(
condition = "input.analysis_type == 'Risk Ratio Forest Plot'",
plotOutput("forest_plot", height = "600px")
)
)
)
)
)
server <- function(input, output) {
# Filtered dataset based on user inputs
filtered_data <- reactive({
data <- adae_data
if (!is.null(input$severity_filter)) {
data <- data %>% filter(AESEV %in% input$severity_filter)
}
if (input$serious_only) {
data <- data %>% filter(AESER == "Y")
}
data
})
# AE Summary Table
output$ae_table <- renderDT({
filtered_data() %>%
group_by(AEDECOD, TRT01P) %>%
summarise(
n = n(),
n_serious = sum(AESER == "Y"),
.groups = "drop"
) %>%
pivot_wider(
names_from = TRT01P,
values_from = c(n, n_serious),
values_fill = 0
) %>%
datatable(options = list(pageLength = 10))
})
# Enhanced AE by Severity Plot
output$severity_plot <- renderPlot({
filtered_data() %>%
ggplot(aes(x = AEDECOD, fill = AESEV)) +
geom_bar(position = "fill") +
facet_wrap(~TRT01P) +
theme_minimal() +
theme(
axis.text.x = element_text(angle = 45, hjust = 1),
plot.title = element_text(hjust = 0.5, face = "bold"),
legend.position = "bottom"
) +
labs(
title = "Adverse Event Severity Distribution by Treatment",
x = "Adverse Event",
y = "Proportion",
fill = "Severity"
) +
scale_fill_viridis_d() +
scale_y_continuous(labels = percent)
})
# Enhanced Time to Onset Plot
output$onset_plot <- renderPlot({
filtered_data() %>%
ggplot(aes(x = ASTDY, fill = TRT01P)) +
geom_density(alpha = 0.7) +
facet_wrap(~AEDECOD) +
theme_minimal() +
theme(
plot.title = element_text(hjust = 0.5, face = "bold"),
legend.position = "bottom"
) +
labs(
title = "Time to Onset Distribution by Adverse Event",
x = "Study Day of Onset",
y = "Density",
fill = "Treatment"
) +
scale_fill_viridis_d()
})
# New Heatmap Plot
output$heatmap_plot <- renderPlot({
event_counts <- filtered_data() %>%
group_by(AEDECOD, TRT01P, AESEV) %>%
summarise(count = n(), .groups = "drop")
ggplot(event_counts, aes(x = AEDECOD, y = AESEV, fill = count)) +
geom_tile() +
facet_wrap(~TRT01P) +
scale_fill_viridis() +
theme_minimal() +
theme(
axis.text.x = element_text(angle = 45, hjust = 1),
plot.title = element_text(hjust = 0.5, face = "bold")
) +
labs(
title = "Adverse Event Frequency Heatmap",
x = "Adverse Event",
y = "Severity",
fill = "Count"
)
})
# New Forest Plot
output$forest_plot <- renderPlot({
# Calculate risk ratios
risk_ratios <- filtered_data() %>%
group_by(AEDECOD, TRT01P) %>%
summarise(n = n(), .groups = "drop") %>%
pivot_wider(names_from = TRT01P, values_from = n, values_fill = 0) %>%
mutate(
risk_ratio = (`Drug A` / sum(`Drug A`)) / (Placebo / sum(Placebo)),
log_rr = log(risk_ratio),
se = sqrt(1/`Drug A` + 1/Placebo),
lower = exp(log_rr - 1.96 * se),
upper = exp(log_rr + 1.96 * se)
)
ggplot(risk_ratios, aes(y = reorder(AEDECOD, risk_ratio))) +
geom_vline(xintercept = 1, linetype = "dashed", color = "gray50") +
geom_errorbarh(aes(xmin = lower, xmax = upper), height = 0.2) +
geom_point(aes(x = risk_ratio), size = 3, color = "darkblue") +
scale_x_log10() +
theme_minimal() +
theme(
plot.title = element_text(hjust = 0.5, face = "bold")
) +
labs(
title = "Risk Ratios of Adverse Events (Drug A vs Placebo)",
x = "Risk Ratio (log scale)",
y = "Adverse Event"
)
})
}
shinyApp(ui, server)I’ve made several enhancements to the visualizations:
Added a Heatmap visualization that shows the frequency of adverse events by severity and treatment group using a color gradient.
Added a Risk Ratio Forest Plot that compares the relative risks of adverse events between treatment groups.
Enhanced the existing plots:
The Severity Plot now shows proportions instead of counts
Added better color schemes using viridis
Improved titles and labels
Enhanced themes and layouts
Added better formatting and styling
Improved the overall UI:
Added a modern theme using bslib's flatly theme
Increased plot sizes for better visibility
Improved layout and spacingThe new visualizations provide different ways to understand the adverse event data:
The heatmap gives a quick visual overview of where the concentrations of adverse events are
The forest plot helps identify which adverse events might be more associated with the treatment
The enhanced severity and onset plots provide clearer comparisons between groupsExport and build
Now that I was satisfied with my prototype app, I exported the generated code. The Shiny Assistant offers a few options for exporting a project in the Shinylive viewer toolbar. I downloaded all the project files and moved them to a project in RStudio.

Note
In addition to downloading the source code, Shiny Assistant provides browser links to the apps, making it easy to share them with others regardless of whether they have R installed. For example, here is the app in editor mode and here is the app in viewer mode.
Clinical trial data tends to follow standardized formats, making it easy to replace the sample data with my actual dataset, stored in adae_data.Rds.
# Generate sample ADAE data
set.seed(123)
n_subjects <- 100
TRT01A <- c("Drug A", "Placebo")
AESEV <- c("Mild", "Moderate", "Severe")
AESEQ <- c("Headache", "Nausea", "Dizziness", "Fatigue", "Rash")
# First, create a vector of how many AEs each subject will have
ae_counts <- sample(1:3, n_subjects, replace = TRUE)
total_records <- sum(ae_counts)
adae_data <- data.frame(
USUBJID = rep(paste0("SUBJ", sprintf("%03d", 1:n_subjects)), ae_counts),
TRT01P = sample(TRT01A, total_records, replace = TRUE),
AEDECOD = sample(AESEQ, total_records, replace = TRUE),
AESEV = sample(AESEV, total_records, replace = TRUE,
prob = c(0.6, 0.3, 0.1)),
ASTDY = sample(1:100, total_records, replace = TRUE),
AESER = sample(c("Y", "N"), total_records, replace = TRUE,
prob = c(0.1, 0.9))
)adae_data <- readRDS("adae.Rds")
n_subjects <- nrow(adae_data)In the Shiny app UI code, I replaced specific variables, such as updating the choices vector to reference columns in adae_data, instead. For example:
choices = AESEVchoices = unique(adae_data$AESEV)Finally, I also swapped all instances of Drug A and Placebo with the corresponding variables from my dataset: A: Drug X and B: Placebo. Midway through, I realized I could have asked the Shiny Assistant to make these changes instead of making them myself.
It sounds incredible, but that’s all I had to do. With these adjustments, I had a fully functional prototype visualizing my ADAE data made with minimal manual coding. See the deployed app on Posit Connect Cloud.
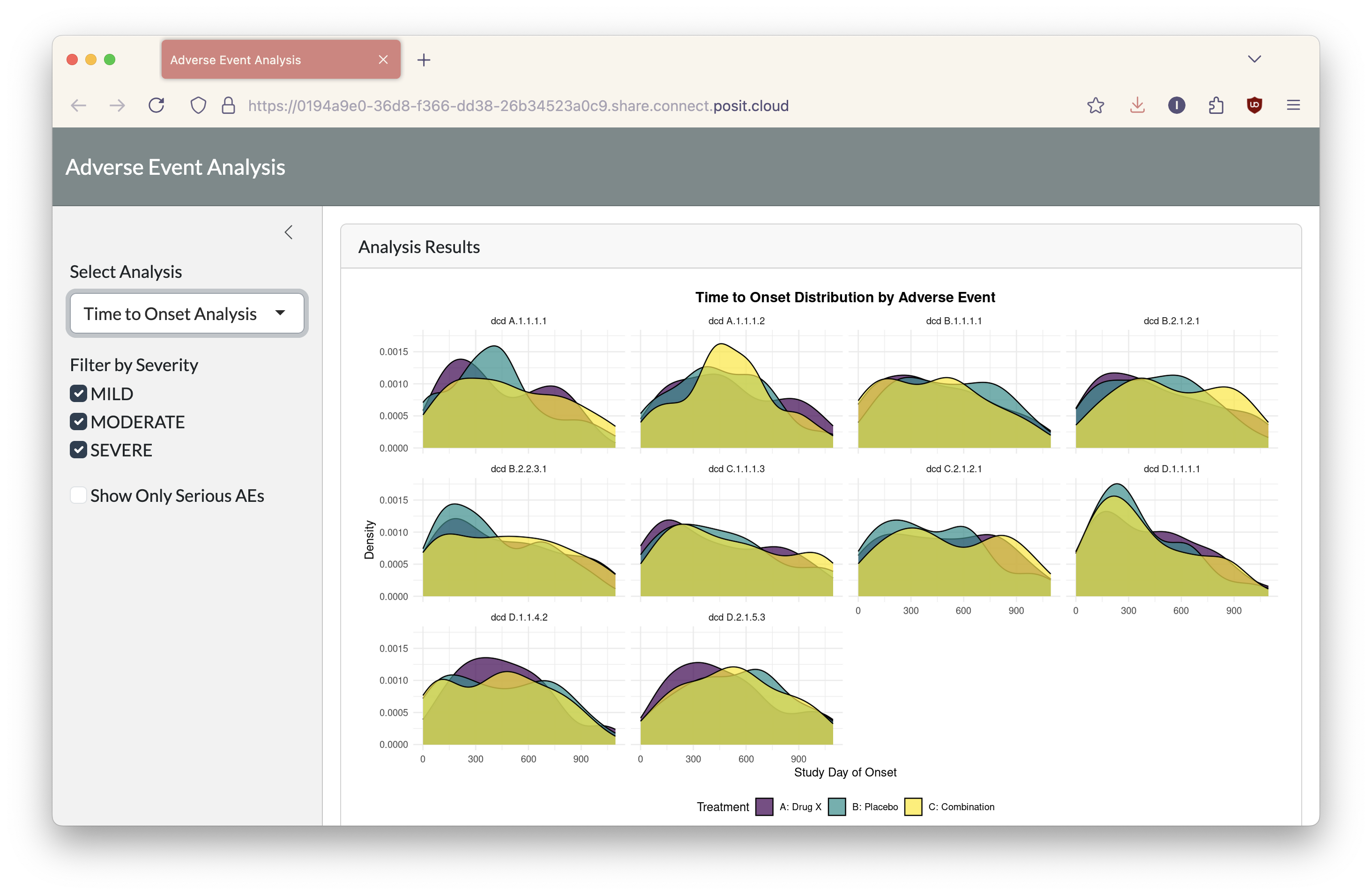
Of course, this is where the human element comes in. I can now refine the dashboard and identify calculation errors in the provided code. While the work was not complete, the Shiny Assistant ensures I had a compelling framework to build upon in a few minutes.
The future of AI-driven app development in pharma
As Joe Cheng stated at R/Pharma, “Summer is coming”. Within life sciences, Shiny Assistant is truly the first of its kind to offer an end-to-end development experience of creating a Shiny application with a chat interface, all presented with a clean user experience. One of the common criticisms of Shiny from those getting started with R programming is overcoming the learning curve to create a prototype application, especially coming from a different statistical programming language such as SAS. Shiny Assistant provides an excellent way to accelerate creating that first prototype. Feedback from early adopters working in pharmaceutical companies has been positive, and they see great potential for the tool to grow further:
- Integration with additional data sources common to a clinical study.
- Fine-tuned knowledge of specific Shiny examples or frameworks to draw upon, such as creating applications with Teal.
- Alternative capability to create customized agents using a similar approach for testing incremental updates to the application code.
Shiny Assistant opens the door for upskilling in Shiny app development, empowering individuals regardless of their R experience to build interactive tools. Try it out today!
Many thanks to Eric Nantz for reviewing this post!



